Amplicon sequencing is a type of DNA sequencing technique commonly used to analyze a specific genetic region. This technique involves the amplification of a DNA fragment, or amplicon, using techniques such as PCR (polymerase chain reaction) to generate a large number of copies. Once these copies are made, they are sequenced to identify genetic variations or mutations. Amplicon sequencing is often used in research and clinical settings due to its high sensitivity and capability to analyze multiple specific regions simultaneously.
Principles of Amplicon Sequencing
Target Selection
The first step in amplicon sequencing is to identify and select the specific region of DNA to be sequenced. This region is called the target. The selection of the target depends on the purpose of the study. For instance, researchers studying a specific genetic disorder may choose a region known to harbor a mutation associated with that disorder.
Amplification
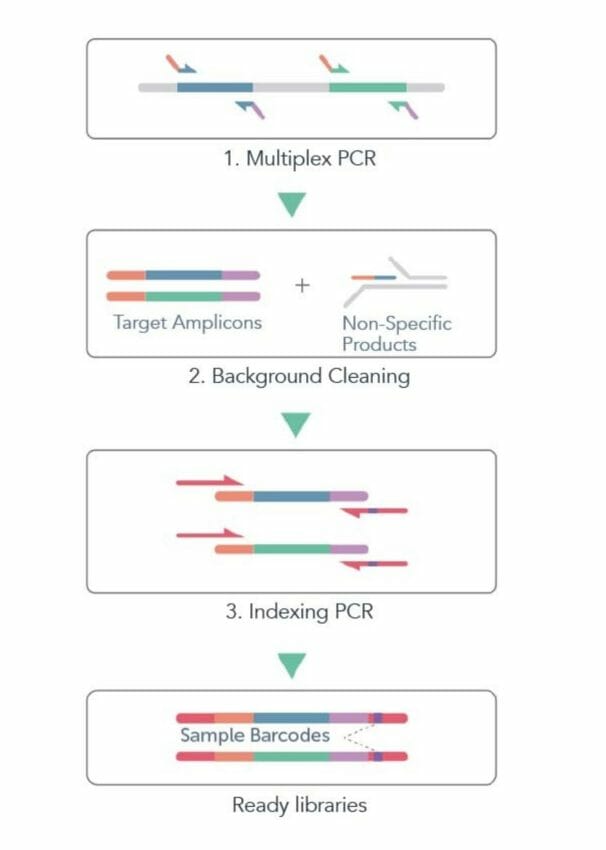
Once the target is selected, the next step is the amplification of the DNA fragment through polymerase chain reaction (PCR). This involves using specific primers that bind to the start and end of the target region. The PCR process generates millions of copies of the target DNA, making it easier to sequence.
Library Preparation
After the target is amplified, a DNA library must be prepared before sequencing. This involves fragmenting the DNA and attaching special adapters to each piece. These adapters enable the amplification of the fragments during sequencing.
Sequencing
After amplification, the DNA samples are sequenced using next-generation sequencing technologies. These technologies allow for high-throughput sequencing, which means that millions of DNA fragments can be sequenced simultaneously.
Data Analysis
The final stage of amplicon sequencing involves analyzing the sequencing data. This includes identifying genetic variations or mutations in the DNA samples. Bioinformatics software tools are often used to assist with this analysis, providing detailed information about the sequence and structure of the target DNA.
Interpretation
After the sequence is analyzed, the results must be interpreted. This involves looking at the data to identify any genetic variations or mutations that are present in the sample. This information can then be used to inform further research or clinical decisions related to diagnosis and treatment of genetic disorders.
Reporting and Visualization
Once the data has been interpreted, it must be reported and visualized in a format that is understandable to researchers, clinicians, and the general public. This can involve creating graphs or tables to represent the sequencing results. Additionally, reports can be generated to provide an overview of the analysis and interpretation of the sequence data.
These principles form the core of amplicon sequencing, allowing researchers and clinicians to analyze specific regions of DNA with high sensitivity and precision.
Applications of Amplicon Sequencing
Amplicon sequencing, also known as targeted or amplicon-based sequencing, has a wide range of applications across various fields of biology and research due to its ability to selectively sequence specific DNA or RNA regions of interest. Some of the key applications of amplicon sequencing include:
Clinical Diagnostics and Personalized Medicine
Amplicon sequencing plays a crucial role in diagnosing and treating genetic diseases. It’s used to identify genetic mutations associated with specific diseases, aiding in early detection and personalized treatment plans based on an individual’s genetic profile.
Genetic Variation and Mutation Analysis
Amplicon sequencing is also used to identify genetic variation and mutations associated with various traits or diseases. This can help researchers gain insight into the underlying molecular basis of these conditions and develop more targeted treatments.
Microbiome Studies
In microbiome research, amplicon sequencing is often employed to study the microbial communities present in an environment, be it the human gut or soil. Researchers sequence a specific region of rRNA gene, which serves as a “barcode” to identify different microbial species.
Population Genetics
Amplicon sequencing has been instrumental in population genetics, where it helps in studying genetic variation within a population. This aids in understanding the evolution, migration patterns, and adaptation strategies of species.
Environmental Monitoring
In environmental science, amplicon sequencing is used to monitor biodiversity, track invasive species, or assess the impact of environmental changes on ecosystems.
Cancer Research
In oncology, amplicon sequencing is used to identify and monitor genetic mutations in tumor cells. This helps in understanding the genetic basis of cancer progression and response to treatment.
Forensic Genetics
Amplicon sequencing is also used in forensic biology, where it helps to identify individuals based on their genetic profiles. It’s often employed for paternity testing and criminal investigations, among other applications.
Viral Genomics
Amplicon sequencing has been used to study the genetic diversity of viruses, such as HIV. This helps in understanding how a virus evolves over time and can aid in developing antiviral treatments or vaccines.
Amplicon sequencing’s ability to focus on specific genomic regions or markers makes it a powerful and cost-effective tool for a wide range of applications, allowing researchers to answer specific biological questions with precision and efficiency.
Recent Advances and Future Directions
In recent years, advancements in amplicon sequencing have greatly improved the speed, accuracy, and affordability of this technique, thereby broadening its scope of application. High-throughput sequencing technologies have dramatically increased the number of samples that can be analyzed simultaneously, improving efficiency in fields such as clinical diagnostics and microbiome research. Furthermore, advances in bioinformatics tools and software have enhanced data analysis capabilities, enabling more precise detection and interpretation of genetic variations or mutations.
Looking forward, the future of amplicon sequencing appears promising. With continued advancements in technology and data analysis, amplicon sequencing is poised to revolutionize personalized medicine by facilitating a more nuanced understanding of an individual’s genetic profile. In addition, advancements in long-read amplicon sequencing are anticipated, which will provide more comprehensive genomic information and enable the sequencing of larger, more complex regions of DNA.
In the environmental realm, amplicon sequencing could prove instrumental in biodiversity conservation efforts by providing insights into species adaptation and evolution in response to environmental changes. On the research front, it is expected that amplicon sequencing will be increasingly employed in multi-omics studies, integrating data from genomics, transcriptomics, metabolomics, and other “omics” to provide a more holistic understanding of biological systems.